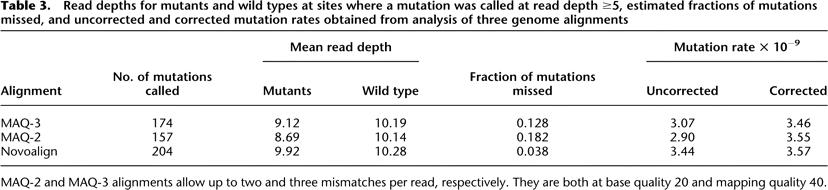
Table 3.
Read depths for mutants and wild types at sites where a mutation was called at read depth ≥5, estimated fractions of mutations missed, and uncorrected and corrected mutation rates obtained from analysis of three genome alignments
Click on table to view larger version.
-
MAQ-2 and MAQ-3 alignments allow up to two and three mismatches per read, respectively. They are both at base quality 20 and mapping quality 40.











