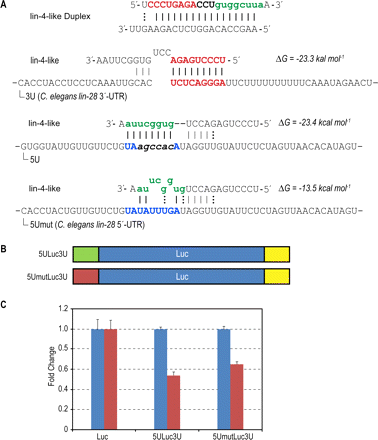
Effect of 5′-UTR interaction site with lin-4-like on reporter expression levels. (A) Predicted interactions between lin-4-like and lin-28-like UTR sequences. The functional strand of the lin-4-like contains an intact cel-lin-4 seed region (red) while the 3′-end is modified (green). There is an extended seed match between the 5′-end of lin-4-like and the wild-type lin-28 3′-UTR binding site (bold red). The 3′-end of lin-4-like is complementary to the artificial lin-28-like 5′-UTR binding site created by introducing a few GC base pairs (bold italics) to form a perfect match. The wild-type lin-28 5′-UTR presents an imperfect match (bold blue). Structure and energy calculations were carried out using RNAhybrid. (B) Schematic showing vector constructs containing firefly luciferase reporter gene used in transfection experiments. lin-28-like 5′-UTR segments containing 8-mer perfectly matched and mutated sites are indicated as 5ULuc3U and 5UmutLuc3U, respectively. (C) Fold changes of Renilla-normalized firefly luciferase expression levels upon co-transfection with lin-4-like (red bars) with respect to nonspecific hsa-miR-16 (blue bars). Error bars represent standard deviation recorded from eight pooled replicates.











