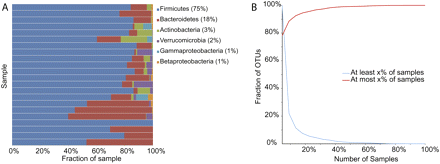
(A) Phylum-level abundance and (B) shared “species” (represented here as 97% OTUs, approximately species level) in 22 human gut samples with depth of coverage of at least 350 sequences per individual. These data are taken from a meta-analysis (Ley et al. 2008a) covering several large Sanger-sequencing studies of humans in different populations (Suau et al. 1999; Hayashi et al. 2002a,b, 2003; Eckburg et al. 2005; Ley et al. 2006c; Nagashima et al. 2006). Interestingly, the results are very consistent with results from both Sanger sequencing and pyrosequencing within a North American population of lean and obese twins (Turnbaugh et al. 2009). Note: No species-level OTUs were shared across all samples with 350 sequences per sample; 1813 OTUs were only present in one sample; the total number of OTUs was 2320.











