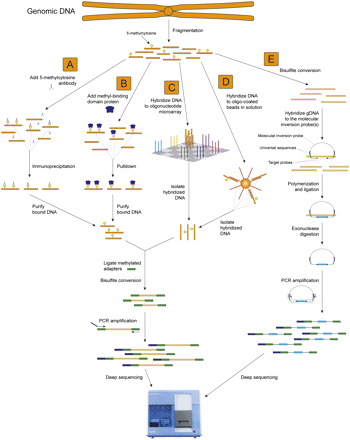
Techniques for enrichment of methylated or target regions prior to BS sequencing. Five approaches that may be used to reduce the complexity of a sample before BS conversion and next-generation sequencing are depicted, targeting methylated regions or select target sequences. (A) MeDIP. Methylated fragments of genomic DNA are immunoprecipitated with an anti-5-methylcytosine antibody. Purified, immunoprecipitated DNA is ligated to double-stranded universal adapter sequences in which all cytosines are methylated. Sodium bisulfite treatment converts unmethylated cytosines to thymine, after which library yield enrichment by PCR with primers complementary to the universal adapter sequences produces the final library that can be sequenced. (B) MBD. Methylated fragments of genomic DNA are isolated from a complex mix of fragmented genomic DNA with a methyl binding domain protein, after which adapter ligation, BS conversion, and PCR enrichment are performed as in A. (C) Microarray capture. Target sequences within a complex mix of fragmented genomic DNA are captured by hybridization to specific oligonucleotides on the surface of a microarray. Following isolation of the hybridized genomic DNA, adapter ligation, BS conversion, and PCR enrichment are performed as in A. (D) Capture in solution. Specific target regions within a mix of fragmented genomic DNA are captured by hybridization to specific oligonucleotides attached to beads in solution. Following isolation of the hybridized genomic DNA, adapter ligation, BS conversion, and PCR enrichment are performed as in A. (E) Molecular inversion probe capture. Fragmented genomic DNA is BS converted, after which molecular inversion probes are added that are designed to hybridize to specific target sequences after conversion. Polymerization primed by the 3′ end of the molecular inversion probe followed by ligation generates a circular molecule that contains the target sequence and is not digested by subsequent exonuclease treatment. PCR using primers that hybridize to the ends of the molecular inversion probes allows amplification of the target region, to which double-stranded universal adapter sequences are ligated to produce a library that is sufficient for next-generation sequencing.











