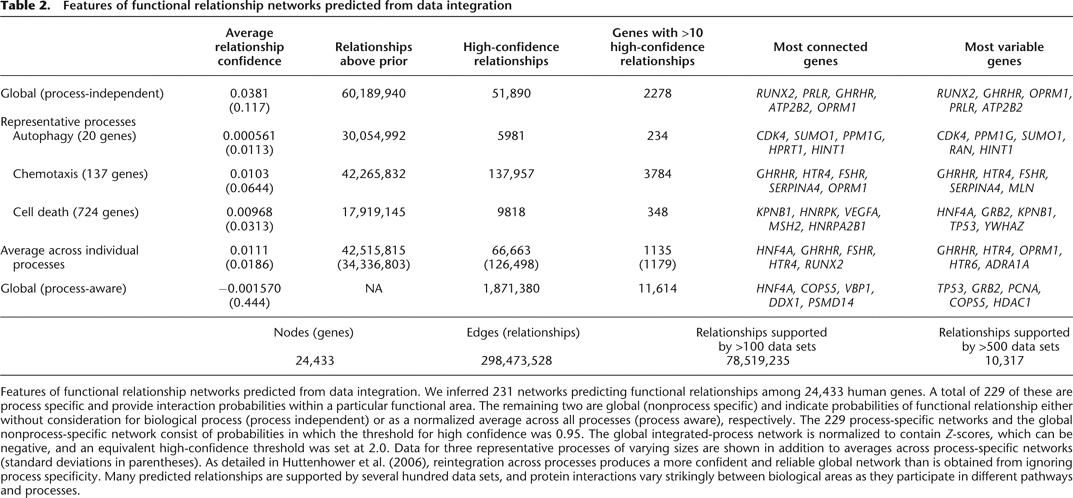
Features of functional relationship networks predicted from data integration
Click on table to view larger version.
-
Features of functional relationship networks predicted from data integration. We inferred 231 networks predicting functional relationships among 24,433 human genes. A total of 229 of these are process specific and provide interaction probabilities within a particular functional area. The remaining two are global (nonprocess specific) and indicate probabilities of functional relationship either without consideration for biological process (process independent) or as a normalized average across all processes (process aware), respectively. The 229 process-specific networks and the global nonprocess-specific network consist of probabilities in which the threshold for high confidence was 0.95. The global integrated-process network is normalized to contain Z-scores, which can be negative, and an equivalent high-confidence threshold was set at 2.0. Data for three representative processes of varying sizes are shown in addition to averages across process-specific networks (standard deviations in parentheses). As detailed in Huttenhower et al. (2006), reintegration across processes produces a more confident and reliable global network than is obtained from ignoring process specificity. Many predicted relationships are supported by several hundred data sets, and protein interactions vary strikingly between biological areas as they participate in different pathways and processes.











