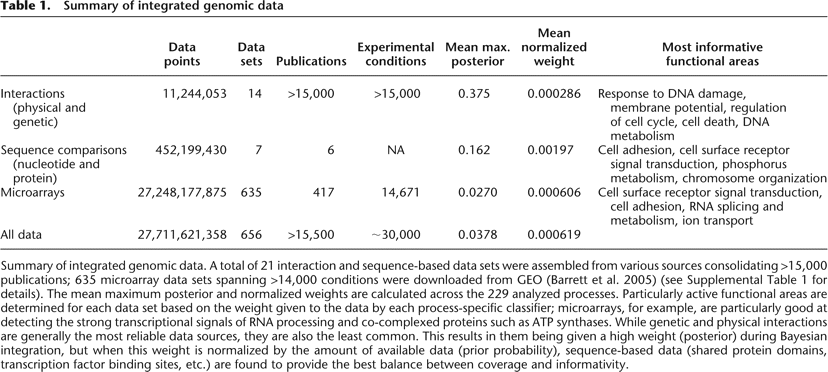
Summary of integrated genomic data
Click on table to view larger version.
-
Summary of integrated genomic data. A total of 21 interaction and sequence-based data sets were assembled from various sources consolidating >15,000 publications; 635 microarray data sets spanning >14,000 conditions were downloaded from GEO (Barrett et al. 2005) (see Supplemental Table 1 for details). The mean maximum posterior and normalized weights are calculated across the 229 analyzed processes. Particularly active functional areas are determined for each data set based on the weight given to the data by each process-specific classifier; microarrays, for example, are particularly good at detecting the strong transcriptional signals of RNA processing and co-complexed proteins such as ATP synthases. While genetic and physical interactions are generally the most reliable data sources, they are also the least common. This results in them being given a high weight (posterior) during Bayesian integration, but when this weight is normalized by the amount of available data (prior probability), sequence-based data (shared protein domains, transcription factor binding sites, etc.) are found to provide the best balance between coverage and informativity.











