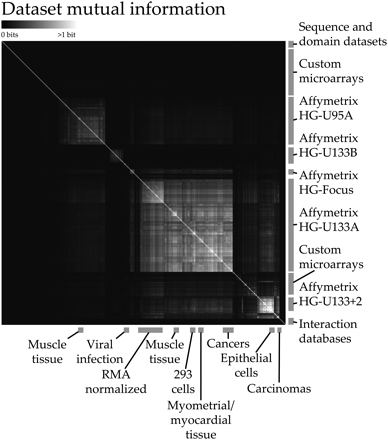
Overview of hierarchically clustered mutual information (MI) between genomic data sets. We used MI among 656 genomic data sets to perform regularization of the parameters of our 230 process-specific Bayesian classifiers. Data sets with a greater proportion of shared information were more heavily mixed with a uniform prior, resulting in the overall up-weighting of particularly unique and informative data. Additionally, a global view of the mutual information scores reveals structure in the data. Primarily platform-based effects can be observed among the expression data sets we obtained from GEO (Barrett et al. 2005), most of which use Affymetrix arrays; tissue type, cell type, and array normalization algorithms can all cause small amounts of information to be shared between many data sets. For example, Robust MultiArray (RMA) normalization causes a noticeable shift in the information shared among HG-U133A arrays. While the amount of MI between any two data sets is generally low (this figure saturates at one bit of shared information), an accumulation of many small overlaps can result in overconfidence during Bayesian data integration, accounting for the success of parameter regularization.











