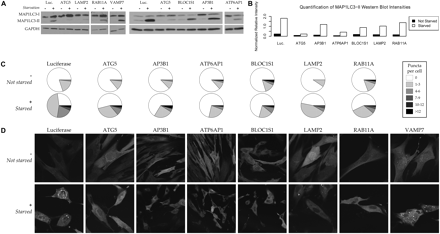
Impaired autophagosome formation confirms the predicted involvement of AP3B1, ATP6AP1, BLOC1S1, LAMP2, and RAB11A in human macroautophagy. Our functional maps predict AP3B1, ATP6AP1, and BLOC1S1 to be involved in autophagy; an early version also predicted the involvement of LAMP2, RAB11A, and VAMP7 in the process, which recycles cellular biomass in order to survive under conditions of starvation or stress. While VAMP7 knockdowns showed no effect (see Discussion), siRNA knockdowns of the other five proteins inhibited normal autophagy. (A) Measurement of the MAP1LC3-I and autophagosome-bound MAP1LC3-II isoforms by immunoblotting. Under a control condition (luciferase siRNA), starvation (+) induces autophagy in human fibroblasts and up-regulates the autophagy marker MAP1LC3-II; this up-regulation is generally inhibited by knockdown of proteins required for autophagy, e.g., ATG5. (B) Quantification of MAP1LC3-II band intensities. Intensities for each condition are calculated relative to GAPDH using the ImageJ software. Replicates (e.g., controls run on multiple gels) have been averaged when available. (C) Quantification of punctate autophagosome formation. The numbers of fluorescent puncta (MAP1LC3-II-labeled autophagosomes) per cell were averaged over counts from three independent investigators in 10 images per normal (−) or starvation (+) condition, unlabeled and randomized (80 images total; see Supplemental Fig. 5 for standard errors). The resulting distribution of puncta frequencies is low under all nonstarved conditions and significantly increased under a negative control (luciferase) condition. It is only slightly increased for the ATG5 positive control and for the AP3B1, ATP6AP1, BLOC1S1, LAMP2, and RAB11A predictions. (D) Punctate localization of fluorescent GFP-LC3 to the autophagosome during autophagy. Under normal conditions (−), MAP1LC3-I is localized diffusely through the cytoplasm; starvation (+) induces autophagy and localization to the autophagosome membrane. Knockdowns of ATG5 (positive control) or the five validated genes abrogate this localization, indicating that these proteins are required for successful macroautophagy.











