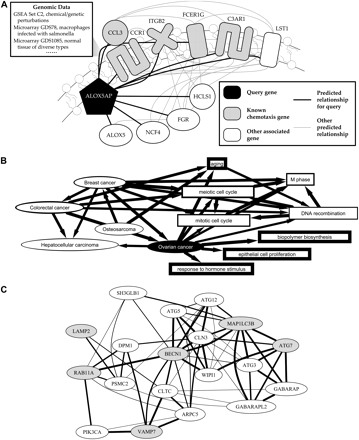
Results of functional mapping from data integration. Functional maps derived from experimental data integration provide information on groups of genes, including cross-talk between pathways, processes, and genes associated with genetic disorders. In all figure parts, thicker edges indicate stronger associations. (A) The process-specific functional relationship networks underlying functional maps can themselves provide information on individual genes' and modules' behavior in the underlying genomic data. Focusing on ALOX5AP, a membrane protein participating in leukotriene synthesis highlights a predicted association with the process of chemotaxis in leukocytes, driven by multiple predicted relationships with known chemotaxis proteins. This represents an instance of functional under-annotation; while ALOX5AP has not been formally cataloged as participating in chemotaxis, its immediate biosynthetic product LTB4 is a known activator of chemotaxis (Peters-Golden and Brock 2003). (B) Associations between genetic disorders and biological processes. To validate functional mapping's ability to discover disease/process associations from data, a focus on ovarian cancer—known to be influenced by at least seven genes (Online Mendelian Inheritance in Man 2008)—we predict associations with the cell cycle, cell proliferation, and hormone stimulus, as well as with several other cancers. These associations are each based on relationships among individual genes predicted from integrated genomic data; directed arrows point to the gene group in which the background connectivity was calculated. As above, additional novel predictions can be explored online using HEFalMp. (C) Visualization of a functional map generated by querying a custom gene set. We chose to focus on the known autophagy proteins ATG7, BECN1, and MAP1LC3B, in addition to genes of interest LAMP2, RAB11A, and VAMP7, in the context of autophagy. This extracts two clear clusters of predicted autophagy-specific functional relationships, one consisting mainly of known autophagy proteins and one enriched for ER/Golgi and vesicular trafficking proteins (including the three test genes). This led us to experimentally test and confirm the hypothesis that LAMP2 and RAB11A (as well as AP3B1, ATP6AP1, and BLOC1S1) are involved in macroautophagy in amino acid-starved human fibroblasts.











