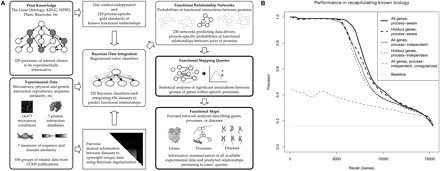
Overview and performance of genomic data integration for functional mapping. (A) Data from ∼30,000 genome-scale experiments (∼15,000 microarray conditions and ∼15,000 interaction and sequence-based assays) were organized into 656 related data sets (Supplemental Table 1). These data sets were used as inputs for 229 process-specific naïve Bayesian classifiers each trained to predict functional relationships specific to a particular biological area and one process-independent global classifier. Mutual information was calculated between each pair of data sets and used to regularize these classifiers and prevent overconfident predictions. Each classifier was used to infer a predicted functional relationship network for a particular biological process. These networks were then analyzed to find statistically significant sets of functional relationships spanning gene groups of interest. This results in functional maps focusing on individual genes, groups of genes, biological processes, or genetic disorders. Each map provides an informative summarization of the genomic data collection focused on the current biological entity of interest. (B) Performance of predicted functional relationship networks in recapitulating known biology. To confirm that the predicted functional relationships underlying our functional maps were accurate, we scored their ability to recover information from a held-out portion (25% of genes) of our gold standard. This evaluation includes the global process-independent network tested on all genes and the holdout set, a process-aware global mean of the process-specific networks tested on all genes and the hold-out set, and an unregularized global process-independent network tested on all genes. Ranking of functionally related gene pairs is performed by comparing predicted probabilities based on data integration with the known relationships in the held-out test set. Results for individual process-specific networks appear in Supplemental Figure 1 and Supplemental Table 3. Precision is well above baseline, and since naïve classifiers are generally robust to overfitting, performance of the hold-out set is only slightly below that of the entire genome. Bayesian regularization provides a large performance increase at low recall by preventing overconfident predictions.











