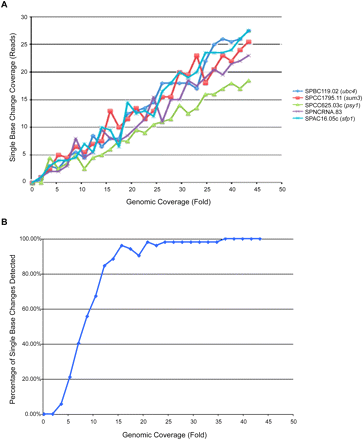
Relationship between library depth and single base change coverage. (A) The 17.04 million Solexa reads derived from swi*603 strain DI36 were sampled randomly to generate 25 libraries ranging from 0.25× to 43× coverage relative to the S. pombe reference genome. For each library, reads were aligned and single base changes detected using MAQ, and the number of sequences spanning five single base changes (including ubc4-G48D) was determined. The relationship between library depth and single base change coverage is generally linear but coverage can range from 1.5- to twofold at a given single base change, presumably due to localized differences in sequencing efficiency. (B) The relationship between DI36 library depth and the percentage confidence of finding all 52 single base changes shared between Broad 972h−, wt DG21, and swi*603 strain DI36. Approximately 22× coverage is required to get to 98% confidence of finding all 52 single base changes.











