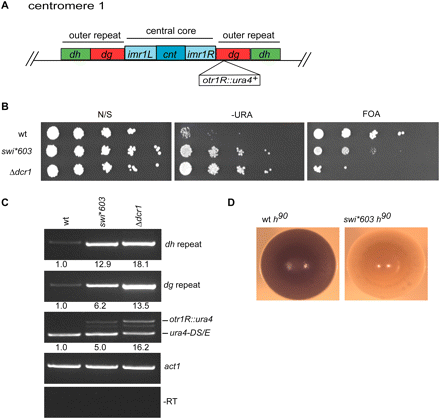
swi*603 is required for centromeric heterochromatin formation. (A) Cartoon of S. pombe centromere 1. The central core domain, where the kintochore forms, is flanked by outer repeat domains composed of dg and dh repeat sequences. The position of the centromeric otr1R∷ura4+ reporter gene used in this study is indicated. (B) swi*603 affects silencing of centromeric heterochromatin. Serial dilution (plating) assays were performed to measure the expression of the pericentromeric otr1R∷ura4+ reporter gene in wt, swi*603, and RNAi-defective Δdcr1 deletion cells. Cells were serially diluted 1/10 (starting with 1 × 104 cells) and spotted onto plates with no selection (N/S), plates lacking uracil (−URA), and counter-selective plates containing 5-fluoro-orotic acid (FOA). Yeast with an active ura4+ gene convert FOA to fluorodeoxyuridine, which is toxic to cells. (C) swi*603 results in the accumulation of transcripts derived from heterochromatic dg and dh centromere repeats. RT-PCR was performed using total RNA isolated from indicated strains to measure the amount of transcript derived from dg and dh centromeric repeats. In each sample, enrichment of each primer pair was measured relative to the control primer pair act1. Centromeric otr1R∷ura4+ expression was also analyzed and compared with the transcriptionally active but nonfunctional mini-gene ura4-DS/E at the ura4+ locus. –RT, minus reverse transcription. (D) swi*603 affects mating-type switching. Iodine staining phenotypes of homothallic (h90) wt strain SPG17 and swi*603 strain. Strains were streaked onto sporulation medium and allowed to grow at 26°C for 5 d before staining with iodine vapors.











