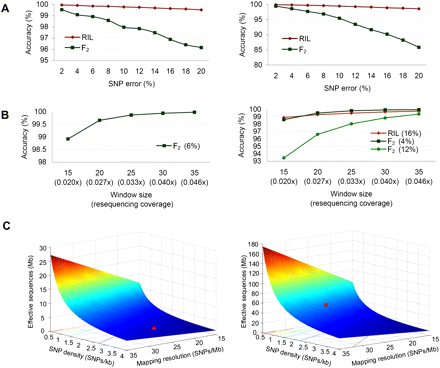
Simulation of genotype calling accuracy. (A) Effect of parental genome sequence quality on calling accuracy. (Left) One parent has high-quality genome sequences that give an SNP error rate of 1%, while the genome sequence quality of the other parent is allowed to vary and gives SNP error rates from 2% to 20%. (Right) Genome sequence qualities of both parents are allowed to vary and give the same SNP error rates from 2% to 20%. Two types of populations, RIL and F2, are considered, with ratios of three genotypes set at 49.5:1:49.5 and 1:2:1, respectively. Window size is set at 15. Genotype calling accuracy is calculated according to Equation 8 in Methods. (B) The effect of window size on calling accuracy. (Left) The critical error rate of 6% that drops the calling accuracy of F2 below 99% in the above figure is used. (Right) Three critical error rates are used, including 16% for both parents that drops the calling accuracy of RIL below 99%, 4% for both parents that drops the accuracy of F2 below 99%, and 12% for both parents that drops the accuracy of F2 below 95%, in the above figure. When window sizes are measured by the number of SNPs covering the same physical distance, increase in window sizes is equivalent to the increase in resequencing coverage. Rice is taken as an example to show resequencing coverage for the corresponding window size. (C) The amount of effective sequences (Se) required for a RIL to reach a range of mapping resolutions (R) as SNP densities (D) vary. (Left) Simulation for the rice genome size, 389 Mb. Red dot indicates the location of the rice RIL of this study (D = 3.2 SNPs/kb, R = 25 SNPs/Mb). (Right) Simulation for the mouse genome size, 2500 Mb. Red dot indicates Se required for a mouse RIL with D = 1.3 and R = 25.











