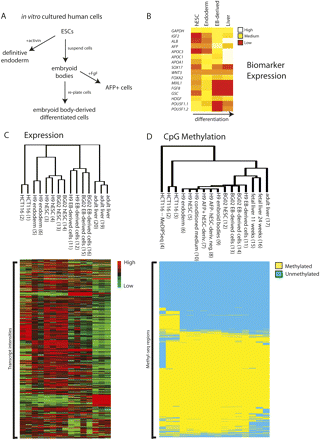
Expression and methylation changes in cells derived from hESCs and human tissues. (A) hESC in vitro differentiation scheme. (B) Expression of key genes in the liver differentiation pathway measured with expression arrays. Expression values are the average of intensities across all biological replicates and both parental hESC lines for each particular tissue. (C) Hierarchal clustering by genes and by samples of log-transformed, normalized expression data for key tissues in the endoderm-lineage model. Shown are a set of 2700 genes that clustered well. Sample numbers are shown in parentheses (see Supplemental Table 1). (D) Hierarchal clustering of the 90,612 Methyl-seq regions for each of the tissues assayed. HCT116 was used in Methyl-seq development and validation and shows reproducibility between biological replicates and concordance with MeDIPSeq data (for more information, see Supplemental Text; Supplemental Fig. 9). Library numbers are shown in parentheses (see Supplemental Table 2).











