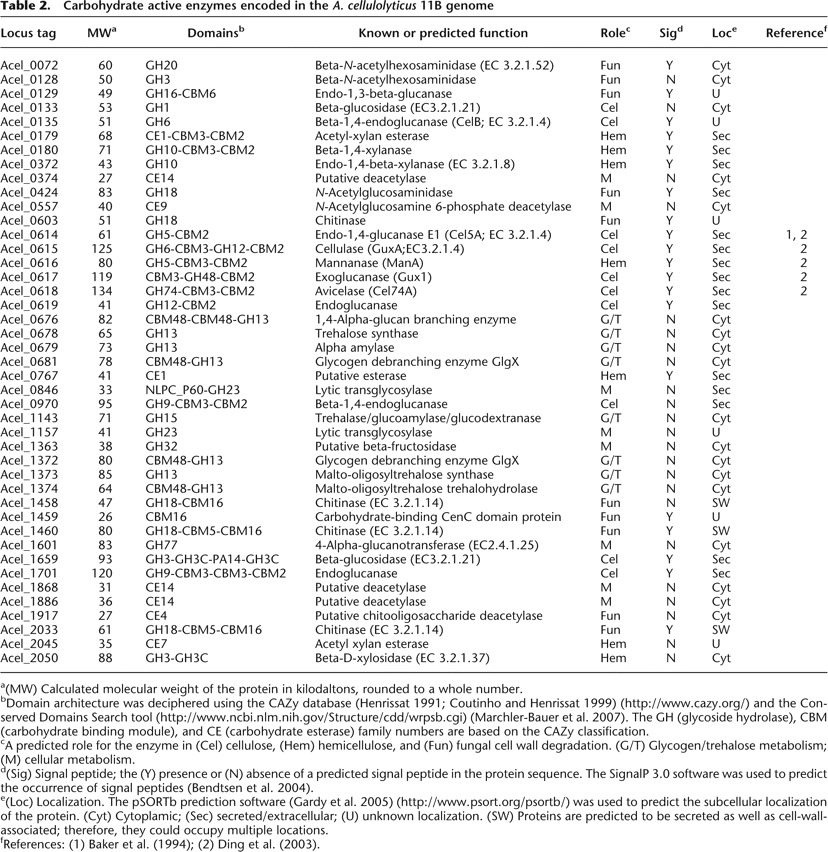
Carbohydrate active enzymes encoded in the A. cellulolyticus 11B genome
Click on table to view larger version.
-
a(MW) Calculated molecular weight of the protein in kilodaltons, rounded to a whole number.
-
bDomain architecture was deciphered using the CAZy database (Henrissat 1991; Coutinho and Henrissat 1999) (http://www.cazy.org/) and the Conserved Domains Search tool (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) (Marchler-Bauer et al. 2007). The GH (glycoside hydrolase), CBM (carbohydrate binding module), and CE (carbohydrate esterase) family numbers are based on the CAZy classification.
-
cA predicted role for the enzyme in (Cel) cellulose, (Hem) hemicellulose, and (Fun) fungal cell wall degradation. (G/T) Glycogen/trehalose metabolism; (M) cellular metabolism.
-
d(Sig) Signal peptide; the (Y) presence or (N) absence of a predicted signal peptide in the protein sequence. The SignalP 3.0 software was used to predict the occurrence of signal peptides (Bendtsen et al. 2004).
-
e(Loc) Localization. The pSORTb prediction software (Gardy et al. 2005) (http://www.psort.org/psortb/) was used to predict the subcellular localization of the protein. (Cyt) Cytoplamic; (Sec) secreted/extracellular; (U) unknown localization. (SW) Proteins are predicted to be secreted as well as cell-wall-associated; therefore, they could occupy multiple locations.
-
fReferences: (1) Baker et al. (1994); (2) Ding et al. (2003).











