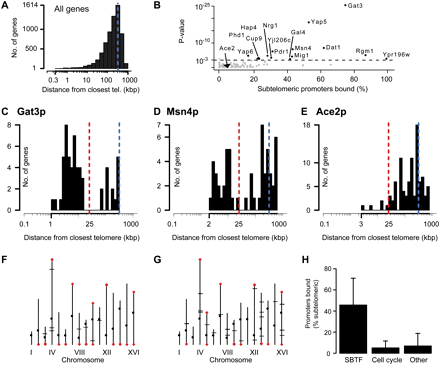
Transcription factors that preferentially bind sequences at subtelomeres. (A) Background telomere distance profile (TDP) for all yeast promoters. (B) Overview of TDPs for rich-medium promoter-binding profiles (Harbison et al. 2004) compared against the background distribution. Each dot represents data for one TF, its significance score on the y-axis (the P-value of a one-sided Kolmogorov-Smirnov test) versus the percentage of bound promoters that are subtelomeric on the x-axis. Subtelomere binding transcription factors (SBTFs) having bimodal TDPs more significant than the P-value threshold of 0.001 are labeled and indicated in black. (C–E) TDPs for Gat3p (C), the statistically most significant SBTF; Msn4p (D), a SBTF that is a master regulator of stress response genes; and Ace2p (E), a cell cycle regulator that is representative of TFs having a TDP similar to the background. Blue broken lines correspond to the blue broken line in A. The red broken line indicates the 25-kb cutoff distance used to categorize subtelomeric genes. (F,G) Plots showing the location of promoters bound on the 16 yeast chromosomes by Gat3p (F) and Msn4p (G). Red diamonds indicate binding events located within 25 kb of a telomere. Black ticks indicate nontelomeric binding events. Small black dots mark the centromere on each chromosome. (H) The average percent of subtelomeric promoters bound by three groups of TFs: SBTFs, cell cycle TFs (those annotated with the “cell cycle” Gene Ontology term GO:0007049), and the remaining TFs. Error bars, SD.











