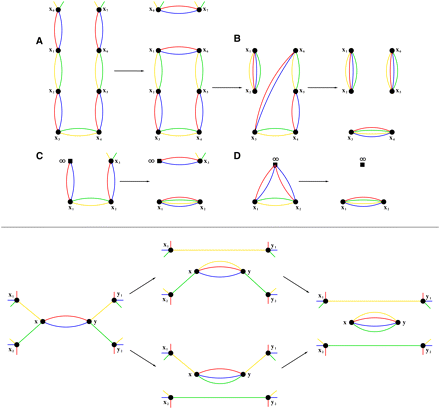
(Top panel) Processing good paths using a 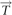 -consistent red–blue multi-color. (A) A good path on vertices x1, x2,…, x6 is transformed into a cycle on the same vertices by extending it into x0, x1, x2,…, x6, x7 and performing a 2-break on the multi-edges (x0, x1) and (x6, x7). (B) Transformation of a good cycle on 6 vertices into complete multi-edges with a 2-break on the multi-edges (x1, x2), (x3, x4) followed by a 2-break on the multi-edges (x1, x4), (x5, x6). (C) A 2-break on an irregular edge corresponds to a reversal involving chromosome ends. (D) A 2-break on two irregular edges corresponds to a fusion. (Bottom panel) Two ways of transforming a fair edge (x, y) into a good edge: (top) by a 2-break on yellow edges or (bottom) by a 2-break on green edges. In either case, the follow-up processing of the generated simple path results in the same graph
with the complete multi-edge (x, y).
-consistent red–blue multi-color. (A) A good path on vertices x1, x2,…, x6 is transformed into a cycle on the same vertices by extending it into x0, x1, x2,…, x6, x7 and performing a 2-break on the multi-edges (x0, x1) and (x6, x7). (B) Transformation of a good cycle on 6 vertices into complete multi-edges with a 2-break on the multi-edges (x1, x2), (x3, x4) followed by a 2-break on the multi-edges (x1, x4), (x5, x6). (C) A 2-break on an irregular edge corresponds to a reversal involving chromosome ends. (D) A 2-break on two irregular edges corresponds to a fusion. (Bottom panel) Two ways of transforming a fair edge (x, y) into a good edge: (top) by a 2-break on yellow edges or (bottom) by a 2-break on green edges. In either case, the follow-up processing of the generated simple path results in the same graph
with the complete multi-edge (x, y).











