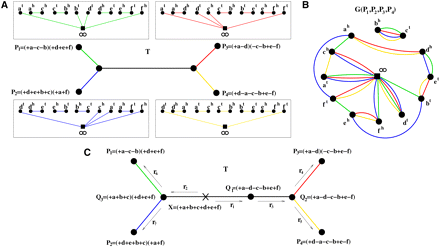
(A) A phylogenetic tree T with four linear genomes P1, P2, P3, P4 (represented as green, blue, red, and yellow graphs, respectively) at the leaves. The obverse edges are not shown. (B) The multiple breakpoint graph G(P1, P2, P3, P4) is a superposition of graphs representing genomes P1, P2, P3, P4. The multi-degrees of regular vertices vary from 1 (e.g., vertex bh) to 3 (e.g., vertex eh). (C) The same phylogenetic tree T with all intermediate genomes specified and a genome X selected as a root. A T-consistent transformation of X into P1, P2, P3, P4 can viewed as a transformation of the quadruple (X, X, X, X) into the quadruple (P1, P2, P3, P4), where a rearrangement at each step is applied to some copies of the same genome in the quadruple. A particular such transformation
takes the following steps: (X, X, X, X) 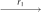 (X, X, Q1, Q1)
(X, X, Q1, Q1) 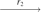 (Q3, Q3, Q1, Q1)
(Q3, Q3, Q1, Q1) 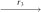 (Q3, Q3, Q2, Q2)
(Q3, Q3, Q2, Q2) 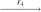 (Q3, Q3, P3, Q2)
(Q3, Q3, P3, Q2)
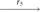 (Q3, Q3, P3, P4)
(Q3, Q3, P3, P4)
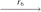 (P1, Q3, P3, P4)
(P1, Q3, P3, P4) 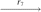 (P1, P2, P3, P4), where r1 is a reversal in two copies of X; r2 is a fission in two copies of X; r3 is a reversal in both copies of Q1; r4 is a fission in one copy of Q2; r5 is a reversal in the other copy of Q2; r6 is a reversal in one copy of Q3; and r7 is a translocation in the other copy of Q3.
(P1, P2, P3, P4), where r1 is a reversal in two copies of X; r2 is a fission in two copies of X; r3 is a reversal in both copies of Q1; r4 is a fission in one copy of Q2; r5 is a reversal in the other copy of Q2; r6 is a reversal in one copy of Q3; and r7 is a translocation in the other copy of Q3.











