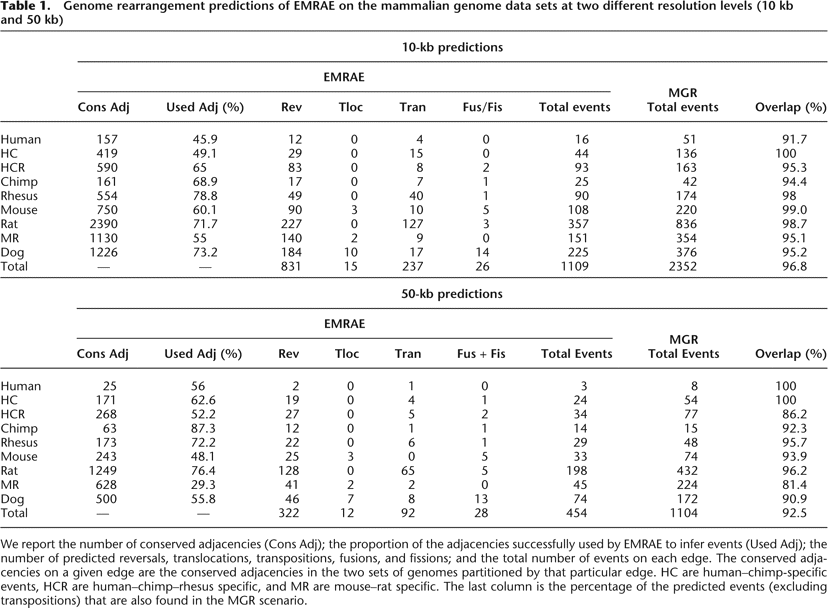
Genome rearrangement predictions of EMRAE on the mammalian genome data sets at two different resolution levels (10 kb and 50 kb)
Click on table to view larger version.
-
We report the number of conserved adjacencies (Cons Adj); the proportion of the adjacencies successfully used by EMRAE to infer events (Used Adj); the number of predicted reversals, translocations, transpositions, fusions, and fissions; and the total number of events on each edge. The conserved adjacencies on a given edge are the conserved adjacencies in the two sets of genomes partitioned by that particular edge. HC are human–chimp-specific events, HCR are human–chimp–rhesus specific, and MR are mouse–rat specific. The last column is the percentage of the predicted events (excluding transpositions) that are also found in the MGR scenario.











