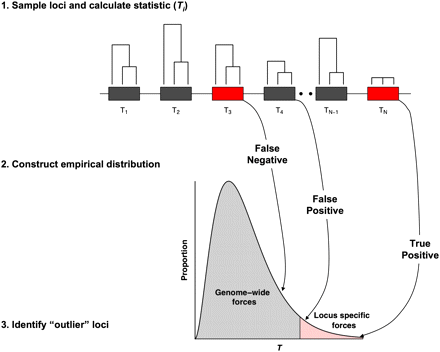
A typical population genomics study design for detecting positive selection. Population genomic studies begin by sampling loci, typically SNPs, throughout the genome. The majority of loci are presumably influenced only by genome-wide forces such as genetic drift (indicated by dark gray boxes). Additional loci, however, may have been subject to locus-specific forces such as selection (indicated by red boxes). Gene genealogies from a sample of three individuals are shown above each locus to emphasize that significant variation in genealogies, and thus, patterns of genetic variation are expected throughout the genome. The extent of variation in genealogies depends on many underlying parameters such as population demographic history and local rates of recombination. For each sampled locus, a statistic of interest (denoted here as Ti for the ith locus) is calculated, an empirical distribution is constructed, and outlier loci are identified in the tail of the empirical distribution. Implicit assumptions of a population genomics approach are that loci are independent, drift influences all loci equally, and selection is strong enough to pull individual loci out into the tail of the empirical distribution. It is important to note that simply occurring in the tail of an empirical distribution does not prove that a locus has been influenced by selection; rather, all one can conclude is that the locus simply has patterns of genetic variation that are unusual in some respect relative to the rest of the genome. Indeed, as shown in the empirical distribution, it is inevitable that some selected loci will not appear as outliers (false negatives) and some neutral loci will appear as outliers (false positives). The lighter red and gray shadings of the empirical distribution reflect that each part of the distribution is a mixture of selected and neutral loci.











