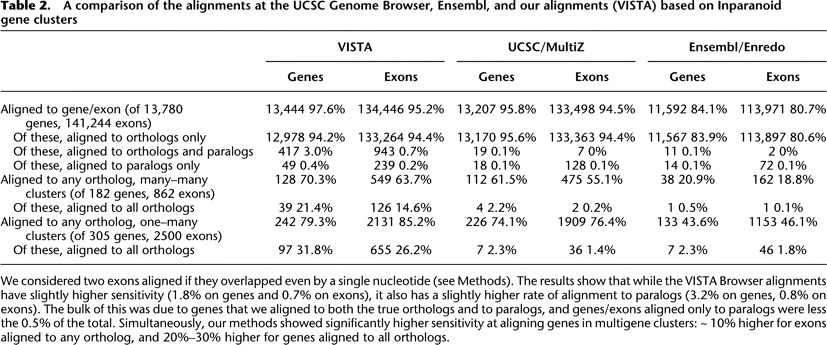
A comparison of the alignments at the UCSC Genome Browser, Ensembl, and our alignments (VISTA) based on Inparanoid gene clusters
Click on table to view larger version.
-
We considered two exons aligned if they overlapped even by a single nucleotide (see Methods). The results show that while the VISTA Browser alignments have slightly higher sensitivity (1.8% on genes and 0.7% on exons), it also has a slightly higher rate of alignment to paralogs (3.2% on genes, 0.8% on exons). The bulk of this was due to genes that we aligned to both the true orthologs and to paralogs, and genes/exons aligned only to paralogs were less the 0.5% of the total. Simultaneously, our methods showed significantly higher sensitivity at aligning genes in multigene clusters: ∼ 10% higher for exons aligned to any ortholog, and 20%–30% higher for genes aligned to all orthologs.











