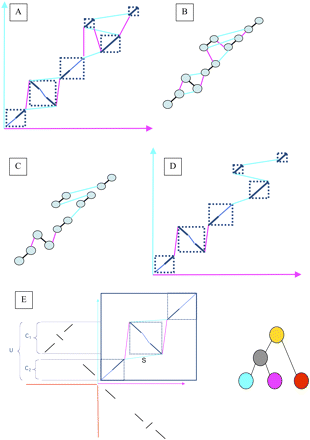
A schematic representation of the reconstruction of ancestral orderings. (A) The result of running SuperMap on a set of local alignments. (B) The corresponding graph representation, with alignment edges colored black, and connection edges colored by the color of the genome in which these syntenic blocks are adjacent. The weight of all of the edges is computed as shown in E. (C) The output of running the maximum matching algorithm: Each node is connected to only one connection edge, as well as the alignment edge. Note that by removing the alignment edges this graph is decomposed into two connected components, which can be solved separately. (D) The translation of the maximum matching output back to the alignments: The result of the algorithm is a chain of alignments, where the letters of the appropriate genome can be inserted between the sequences. These chains can then be used for alignment in higher nodes of the tree. (E) In this example we are recreating the ancestral order of the gray node in the phylogeny on the right. The top right quadrant shows the output of the SuperMap algorithm applied to the blue and purple genomes. The top left and bottom right quadrants show the local hits of the two genomes on the red outgroup. The selected regions on the left are used to compute the score for the blue edge marked S (S = (U − MIN(C 1 ,C 2))/MAX(C 1 ,C 2)). All of the other edges will be scored the same way, and the MWPM problem is solved in the resulting graph. In this particular case the purple genome will have more support for being the ancestral order than the blue genome.











