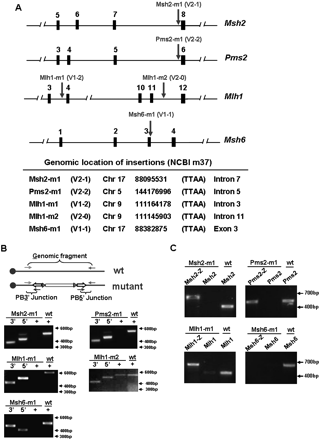
Analysis of 6-TGR clones. (A) Insertion sites of transposons in four DNA mismatch repair genes shown by arrows. The insertion–host junctions were identified by SpPCR and sequence analysis. The vector and reading frames are indicated. (B) Homozygosity analysis of five mutant clones; Msh2-m1, Pms2-m1, Mlh1-m1, Mlh1-m2, and Msh6-m1. Genomic PCR analysis was conducted with primers for the PB3′ and PB5′ transposon/host junctions (mutant alleles, marked as “3′” and “5′,” respectively) or with genomic primers which span the insertion site (wild-type alleles). The wild-type genomic fragments “+” are absent in gene-trap mutant clones but were amplified from Blm-deficient ES cells (marked as “wt”), which indicated that transposon insertions are homozygous in all cases except Mlh1-m2. (C) Wild-type and fusion transcripts in gene-trap mutants. RT-PCR was performed with primers for the upstream exon of the relevant mutated gene to lacZ gene (marked as “Msh2-Z, Pms2-Z, Mlh1-Z, and Msh6-Z”) as well as for the wild-type alleles. The wild-type transcripts are detected in nonmutated Blm-deficient control samples (marked as “wt”) but were absent in all four mutant clones. Fusion transcripts of the expected size were detected in three mutant clones, Msh2-m1, Pms2-m1, and Mlh1-m1.











