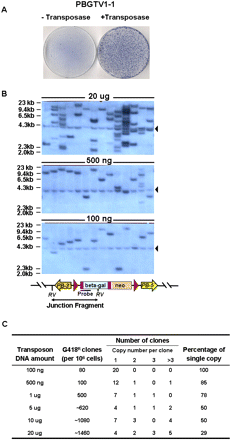
Copy number analysis of PB insertions. (A) PB transposition in the presence of transposase. Five micrograms of transposon gene-trap vector (PBGTV1-1) were electorporated with or without 10 μg of transposase expressing plasmid into 5 × 106 AB2.2 cells. A representative picture of plates obtained from G418 selection. (B) Southern blot analysis of gene-trap clones reveals host/transposon junction fragment in each clone. RV, EcoRV. LacZ, probe, an 800-bp ClaI fragment from pSAgeo (Soriano et al. 1991). Black triangles indicate the endogenous neomycin locus in NGG5-3 ES cells. (C) Summary of copy numbers obtained using different transfection conditions at a single transposase concentration, 20 μg.











