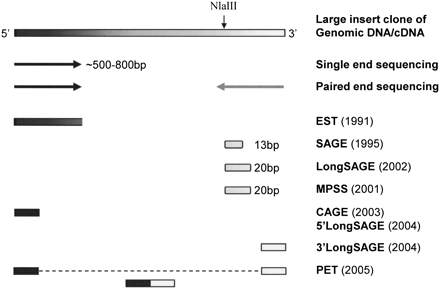
Sequencing-based methods for understanding genetic elements in genomes. The nucleotide information of DNA fragments can be spelled out by complete sequencing analysis. Alternatively, the large size of DNA fragments can be analyzed by single-end sequencing or paired-end sequencing. EST was the first tag-based approach, generating one tag per sequencing read, to represent full-length transcripts of expressed genes. The original SAGE tag extracts 13 bp next to the 3′ most NlaIII site to tag the transcripts. LongSAGE and MPSS use MmeI as the tagging enzyme to generate 20-bp tags that can be specifically aligned to reference genome sequences. The CAGE and 5′ LongSAGE tags are derived from the 5′ end of cDNA fragments, and the 3′ LongSAGE tags are derived from the 3′ end of cDNA fragments. PET covalently combines the 5′ and 3′ signature tags of the same DNA fragment into one ditag unit.











