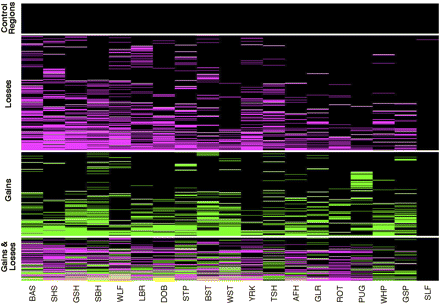
Heatmap representation of CNVs. Each row represents one of the 678 unique CNV regions and columns correspond to dogs. For each CNV region, boxes are colored as black, magenta, and green, depending on whether the individual showed no copy number variation, a loss, or a gain, respectively. CNV regions that show both a loss and a gain within an individual dog (see text) are colored yellow. Horizontal white lines separate CNV calls from single copy control regions, and CNVs that exhibit only losses, gains, or both gains and losses. Within each class, CNV regions are sorted from low to high frequency and from left to right dogs are sorted by decreasing number of CNVs. Dog breeds are abbreviated as follows: Basenji (BAS), Shetland Sheepdog (SHS), German Shepherd (GSH), Siberian Husky (SBH), Wolf (WLF), Labrador Retriever (LBR), Doberman Pinscher (DOB), Standard Poodle (STP), Belgian Shepherd Tervuren (BST), West Highland White Terrier (WST), Yorkshire Terrier (YRK), Boxer (Tasha) (TSH), Afghan Hound (AFH), Golden Retriever (GLR), Rottweiler (ROT), Pug (PUG), Whippet (WHP), German Short Haired Pointer (GSP), and Self-Self hybridization (SLF).











