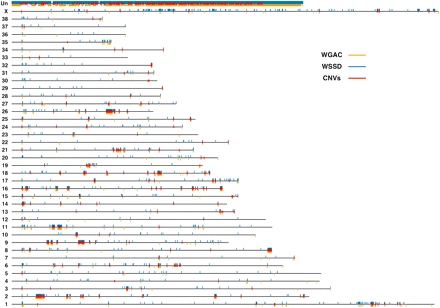
The genomic architecture of canine segmental duplications and CNVs. Black lines represent all 38 canine autosomes, the X chromosome, and the uncharacterized chromosome (Un). Duplicated bases predicted by WGAC and WSSD are plotted as orange and blue rectangles below and above, respectively, each chromosome. Over 80% of chrUn contains duplicated bases. Of the autosomes, chromosomes 9, 16, and 26 possess the highest percentages of duplicated bases (over 4% of each chromosome), while chromosomes 12, 30, and 33 show the least amount of duplicated bases (<0.35% of each chromosome). Unique CNV regions (see text) are denoted by red rectangles.











