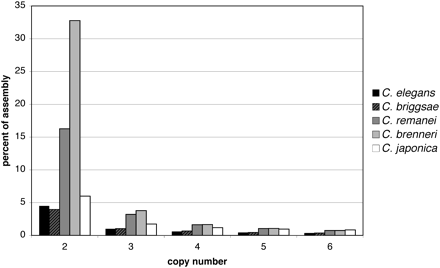
Estimated copy number distributions for five genome assemblies. For each species, a sliding query window of 1000 bp with 500 bp steps was used to identify nonself matches in the assembly. The percentages reported are relative to the size of the total assembly, not to an inferred actual genome size. For the hermaphroditic C. elegans (sequenced by a minimum clone tiling path method) and C. briggsae (sequenced by WGS), all sequences with copy number of two or more likely represent true copy number variation in the genome. For the three gonochoristic species, however, each bin potentially represents a mix between truly paralogous DNA and retained alleles. As the apparent single-copy sequence is at least 55% in all assemblies, the majority of the unrecognized alleles are expected to lie in the two-copy category. Single-copy DNA is not shown.











