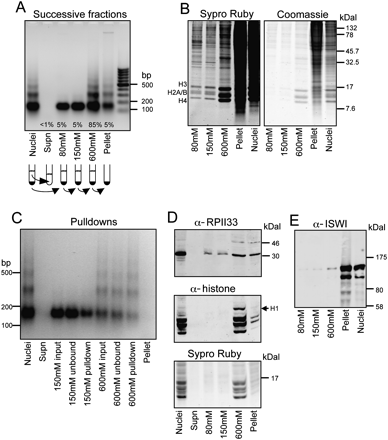
DNA and protein characterization of S2 cell chromatin fractions. (A) Ethidium bromide–stained agarose gel showing a typical MNase ladder of DNA purified from MNase-treated nuclei (Nuclei); followed by ladders of DNA purified from the nuclear supernatant (Supn); successive 80 mM, 150 mM, and 600 mM extractions; and the remaining pellet, as indicated in the cartoon below. The average percentage of total chromatin isolated at each step is indicated. (B) SDS–polyacrylamide gel analysis (SDS-PAGE) of proteins in salt fractions. Equal aliquots of fractions from successive extraction steps in a typical experiment were loaded onto a 16% polyacrylamide gel, which was electrophoresed and stained with Sypro ruby for sensitive detection of histones (left). Restaining with Coomassie blue (right) shows that the pellet includes mostly high-molecular-weight proteins. (C) MNase ladders of 150 mM and 600 mM fractions from a typical pulldown experiment. (D) Western blot analysis of the RPII33 subunit of RNA polymerase II (Pol II) and histones in salt fractions. Equal aliquots of samples were loaded onto a 16% SDS-PAGE gel, electrophoresed, blotted, probed with rabbit RPII33 (Muse et al. 2007) and mouse pan-histone (Roche MAB052) antibodies, followed by IRD700 anti-rabbit and IRD800 anti-mouse secondary antibodies (LI-COR). After blotting, the gel was stained with Sypro ruby. RPII33 is enriched in the insoluble fraction relative to histones. Histone H1 is mostly extracted with the 600 mM fraction as expected (Davie and Saunders 1981). (E) Western blot analysis of the ISWI nucleosome remodeling protein in salt fractions. Equal aliquots of samples were loaded onto a 4%–12% NuPAGE (Invitrogen), gel electrophoresed and blotted to nitrocellulose, and probed with an ISWI antibody (Tsukiyama et al. 1995). Successive extractions release increasing amounts of ISWI, although most of the ISWI (MW 119) that is recovered resides in the insoluble fraction.











