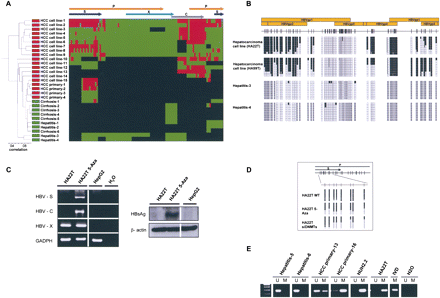
The DNA methylome of HBV. (A) Unsupervised clustering analysis of the HBV DNA methylome in carriers of the virus with pre-malignant lesions such as cirrhosis and hepatitis, primary liver tumors (HCC primary), and liver cancer cell lines (HCC cell line). (Red) Methylated, (green) unmethylated, and (black) deleted sequences. (Top) The HBV genome. (B) Example of bisulfite genomic sequencing analysis of multiple clones for the HBV genome. (Black squares) Methylated and (white squares) unmethylated CpG dinucleotides; (gray squares) deleted genome sequences. (C) DNA methylation-associated silencing of the HBVgp4 (HBV-C) and HBVgp2 (HBV-S) genes in the hepatic carcinoma cell line HA22T and reactivation upon the use of a DNA demethylating agent (5-aza-2′-deoxycytidine). The HBVgp3 remains unmethylated and expressed. (Left) RT-PCRs; (right) Western blots. The hepatoblastoma cell line HepG2, negative for the HBV virus, is used as a negative control. (D) Depletion of DNMT1 and DNMT3B by short interference RNA or upon treatment with the DNA demethylating agent (5-Aza) causes a DNA hypomethylation of the HBV genome in HA22T cells. (E) Methylation-specific PCR analysis of the HBVgp2 sequence in liver tumorigenesis. The presence of a band under the U or M lanes indicates unmethylated or methylated sequences. In vitro methylated DNA (IVD) is shown as a positive control.











