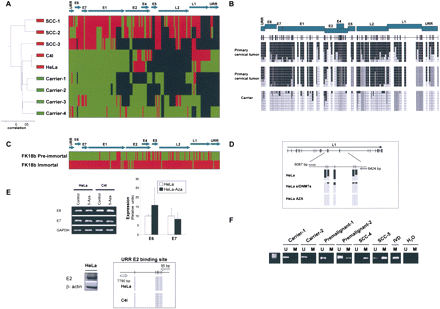
The DNA methylome of HPV18. (A) Unsupervised clustering analysis of the HPV18 DNA methylome in asymptomatic carriers of the virus, primary tumors (SCCs), and cancer cell lines (HeLa and C4I). (Red) Methylated, (green) unmethylated, and (black) deleted sequences. (Top) The HPV18 genome. (B) Example of bisulfite genomic sequencing analysis of multiple clones for the HPV18 genome. (Black squares) Methylated and (white squares) unmethylated CpG dinucleotides; (gray squares) deleted genome sequences. (C) Bisulfite sequencing showing how the mostly unmethylated HPV18 DNA methylome from pre-immortal keratinocytes undergo hypermethylation in immortalized cells. (Red) Methylated and (green) unmethylated sequences. (D) Depletion of DNMT1 and DNMT3B by short interference RNA or upon treatment with the DNA demethylating agent causes a DNA hypomethylation of the HPV18 genome in HeLa cells. (E) The use of a DNA demethylating agent (5-aza-2′-deoxycytidine) does not change the expression of the HPV18 E6 and E7 oncoproteins in HeLa cells in association with an unmethylated URR E2-binding site. (Top) Conventional RT-PCRs and q-RT-PCRs; (bottom) expression of E2 and bisulfite sequencing that demonstrates an unmethylated URR E2-binding site. (F) Methylation-specific PCR analysis of HPV16 E2 sequence in cervical tumorigenesis. The presence of a band under the U or M lanes indicates unmethylated or methylated sequences. In vitro methylated DNA (IVD) is shown as a positive control.











