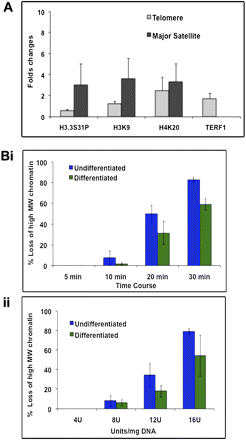
Chromatin immunoprecipitation (ChIP)/PCR analysis and MNase digestion assays. (A) ChIP was performed on undifferentiated and differentiated ES129.1 cells using anti-sera against TERF1, H3.3S31P, H4K20me3, and H3K9me3, followed by real-time PCR analysis using primers specific for telomere or pericentric major satellite DNA. Fold-differences in the enrichment of the ChIP/PCR products (n = 3) were presented as histograms. Following differentiation, H3.3S31P levels at the telomeres was only 0.54-fold (i.e., reduced by 46%) of those in undifferentiated cells, whereas H3K9me3 and H4K20me3 levels at the telomeres were increased by 1.2- and 2.5-fold, respectively. At the pericentric heterochromatin, the levels of H3.3S31P, H3K9me3 and H4K20me3 were increased by 3.0-, 3.6-, and 3.3-fold, respectively. Data for the analysis are shown in Supplemental Table 1. (B) MNase assays (n = 3) were performed on both undifferentiated (i) and differentiated (ii, 6 d) ES129.1 cells by MNase digestion over a time-course (i, 4 units MNase/mg DNA for 5 to 30 min at 37°C) and with various concentrations of MNase (ii, 4–16 units/mg DNA for 5 min at 37°C). Southern blot analysis was performed using radioactive-labeled telomere probe. Change in MNase sensitivity was quantitated by measuring the distance between nucleosomes with the highest and lowest molecular sizes (Gilbert et al. 2007). Results were presented as percentage of loss of high-molecular-weight (MW) chromatin. Telomeric chromatin in differentiated cells was less sensitive to MNase digestion. Examples of individual MNase/Southern analysis are shown in Supplemental Figure 9.











