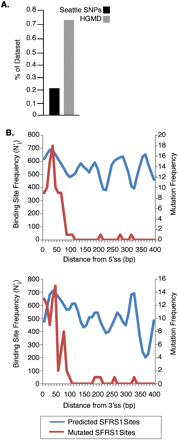
Disruption of SFRS1 binding sites can cause human inherited disease. (A) Single-nucleotide substitutions causing loss of predicted SFRS1 binding sites in the Human Gene Mutation Database (http://www.hgmd.org) and the SeattleSNPs database (http://pga.gs.washington.edu) were identified by scanning reference and mutated exon sequences with the SFRS1 PWM. The proportion of entries in each database giving rise to a loss of SFRS1 sites was then plotted, and a statistically significant difference between the HGMD and SeattleSNPs data sets observed (P < 10−5; Fisher's exact test). (B) Disease mutations resulting in the loss of SFRS1 binding sites were found to be largely confined to exon boundaries. The top and bottom panels plot mutated sites relative to the 5′ and 3′ splice sites, respectively. The blue line in each plot represents the distribution of SFRS1 binding sites throughout the HGMD exons. The red lines correspond to the distribution of sites ablated by disease-causing mutations. These data demonstrate that although binding sites for SFRS1 can be predicted across HGMD exons, disease mutations tend only to disrupt those SFRS1 binding sites that are in close proximity to splice sites.











