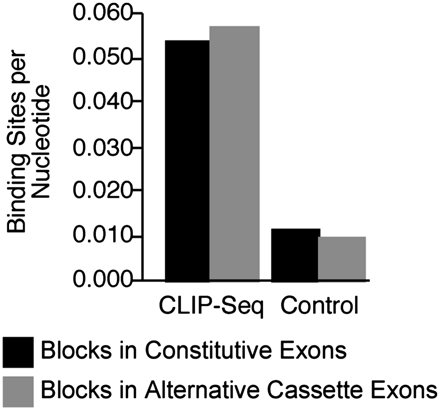
The SFRS1 consensus motif was enriched in blocks identified by CLIP relative to randomly selected blocks from exon sequences. The average number of SFRS1 consensus sites per nucleotide was determined and plotted for sequence blocks in 8693 constitutive (black bars) and 426 alternative cassette exons (gray bars) identified by the CLIP-seq experiment. For the control group, 55 bp regions were selected at random from equal numbers of constitutive or alternative cassette exons not contained in the pool of SFRS1 CLIP-seq data. The Wilcoxon test confirmed the mean binding sites per nucleotide were significantly different for CLIP-seq and control exons (P < 10−22). Binding sites for SFRS1 in alternative cassette exons were found to be modestly enriched relative to constitutive exons (P < 0.005).











