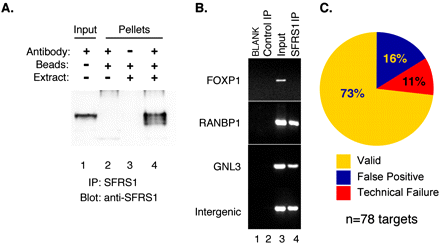
Validation of SFRS1–RNA interactions by RNA–IP RT–PCR. (A) Western blot analysis of proteins precipitated by the anti-SFRS1 monoclonal antibody. SFRS1 was detected in both the input extract (lane 1) and the material immunoprecipitated with anti-SFRS1 (lane 4) but not the control beads (lanes 2,3). The blot was visualized as described in Figure 1. (B) Examples of RT–PCR analysis of endogenous SFRS1–mRNA complexes. RNA extracted from the control IP, input extract, and SFRS1 IP, was reverse transcribed using oligo dT and Superscript III (Invitrogen). A total of 78 different primer sets were used to amplify specific transcripts from cDNA. (C) Summary of RT–PCR validation of 78 randomly selected sequence blocks identified by CLIP-seq. Validated interactions correspond to detectable PCR product in both the input and SFRS1 IP samples. False positive transcripts correspond to PCR products present in the input and the control IP. Technical failures yielded no PCR products from any cDNA sample, indicating that the transcript could not be directly validated under these conditions.











