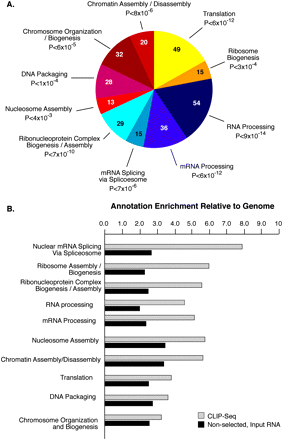
SFRS1 targets are enriched in mRNAs encoding RNA-binding proteins. (A) The top 10 classes of Gene Ontology terms enriched in the CLIP data set relative to the expected ratios in the DAVID database (Hosack et al. 2003). The top 10 classes of targets are all related to gene expression. The numbers of genes observed in each category are indicated in the pie chart. Holm-corrected EASE scores are given for each category (Hosack et al. 2003). (B) Comparison of the top 10 CLIP-enriched Gene Ontology terms with nonselected input mRNA samples. The annotation enrichment relative to the genome is plotted for both CLIP (gray bars) and input (black bars) derived mRNAs. mRNAs encoding splicing factors are most highly enriched in CLIP.











