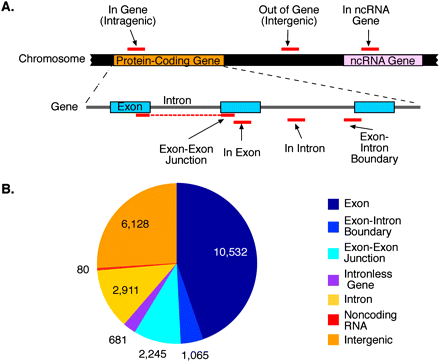
Classification of cis-acting RNA elements bound by SFRS1. (A) Annotation strategy for classifying sequence blocks identified by CLIP-seq. Following alignment of amplicon sequences to the human genome, blocks of overlapping sequences were defined and subsequently annotated using the UCSC Known Gene and Rfam databases. Blocks were classified at the gene level (“In Gene,” “Out of Gene,” and “In ncRNA”) and at the transcript level (“In Exon,” “In Intron,” “Exon–Intron Boundary,” and “Exon–Exon Junction”). None of the blocks presented here overlapped amplicon sequences obtained from input RNA samples. (B) Quantification of block annotations. The majority of blocks were classified as “In Gene” and were predominantly associated with exon sequences. Slightly more than 25% of blocks were defined as intergenic, but binding sites for SFRS1 within these transcripts were found to be highly conserved over evolutionary time (See Supplemental Fig. 2).











