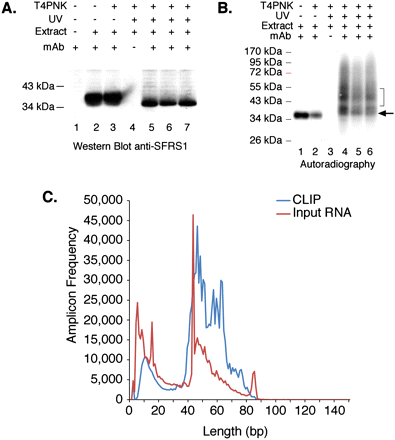
CLIP of SFRS1 from HEK293T cells and amplicon sequencing. (A) Western blot analysis of SFRS1 immunopreciptation (IP) from control and UV-cross-linked cells. CLIP was performed in three independent cultures of HEK293T cells (lanes 5–7). SFRS1 was detected with the same antibody used for IP. The blot was visualized with chemiluminescence (Pierce Super Signal). This panel demonstrates the specificity of the IP; no signal was detected in control IP samples (lane 4), but SFRS1 was efficiently precipitated from both UV-irradiated and control cell extracts. (B) Autoradiograph of 32P-labeled SFRS1–RNA complexes immobilized on nitrocellulose membrane. Following extensive washes of the immunoprecipitated complex, the beads were incubated with T4PNK (NEB) and [γ32P]ATP in order to phosphorylate the 5′ end of RNA fragments. In the absence of UV irradiation, SFRS1 can become phosphorylated. However, this does not require T4PNK (cf. lanes 1 and 2). These data suggest that a subset of SFRS1 is copurified with an SR protein kinase even under these stringent conditions. Importantly, in the absence of UV irradiation, no slower migrating protein–RNA complexes are observed (cf. lanes 1,2 and 4–6). The arrow indicates the position of “free” SFRS1 and the bracket defines the region of nitrocellulose excised from the blot. (C) Histogram comparing the amplicon length distribution with numbers of reads. Amplicons prepared from both CLIP (blue line) and nonselected input RNA samples (red line) were sequenced using the 454 FLX platform and short read reagents. As expected from B, the majority of amplicons (after removal of both linker sequences) are between 40 and 60 bp in length.











