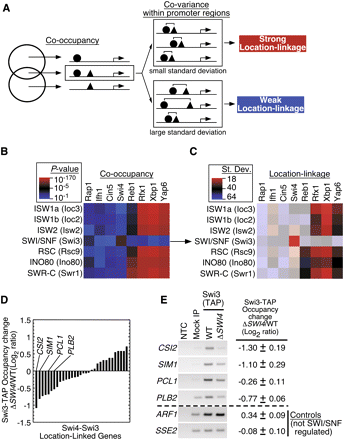
Sequence-specific regulators are location-linked to chromatin remodelers. (A) Schematic overview of the location-linkage analysis. Statistically significant promoter co-occupancy between a sequence-specific regulator and chromatin remodeler was computed using CHITEST function in Excel. Two factors are location-linked if their positions covary at genes in which they co-occupy. The extent to which two factors covary across all co-occupied promoters was determined by computing the standard deviation for the distances between their co-occupied binding locations. (B) Promoter co-occupancy between sequence-specific regulators and chromatin remodelers. Red reflects greater co-occupancy significance; blue, less significance relative to P = 10−5. (C) Location linkage of sequence-specific regulators and chromatin remodelers. Brighter red indicates far less than the average standard deviation of pairwise distances (greater location-linkage). Darker blue indicates greater than average standard deviation (less location-linkage). Co-occupancy pairs that did not pass criterion 1 are shown with lighter shading. (D) ChIP-chip was used to determine the changes in occupancy for Swi3-TAP (SWI/SNF) in a swi4Δ strain. The Swi3-TAP occupancy changes for the genes that are Swi4-Swi3 location linked (panels B,C) are displayed as a bar graph. (E) Standard ChIP was performed on four of the genes most dependent on Swi4 for SWI/SNF recruitment (CSI2, SIM1, PCL1, and PLB2). A no template control (NTC) and a mock IP using an untagged BY4741 strain are shown. The PCR products from four biological replicates were quantified, and the log2 ratios (swi4Δ/WT) ± SD for the Swi3-TAP changes in occupancy are displayed to the right of the PCR product for each gene. Loss of SWI4 did not diminish Swi3-TAP expression or its global distribution (Supplemental Fig. S9).











