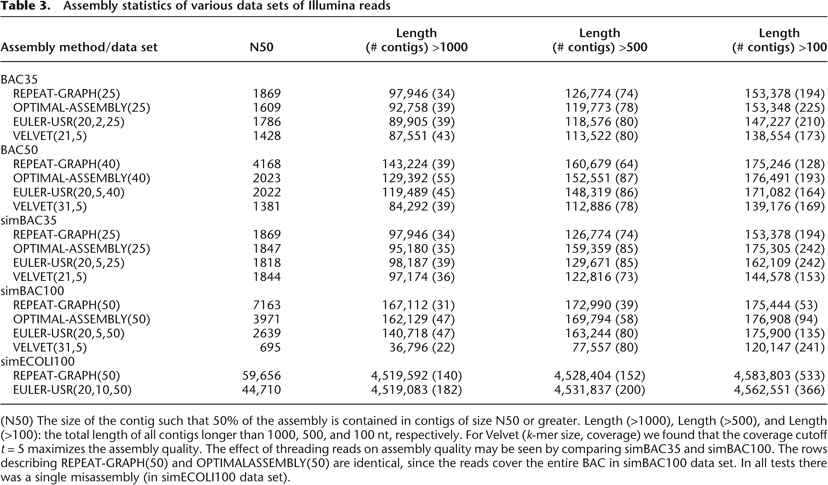
Assembly statistics of various data sets of Illumina reads
Click on table to view larger version.
-
(N50) The size of the contig such that 50% of the assembly is contained in contigs of size N50 or greater. Length (>1000), Length (>500), and Length (>100): the total length of all contigs longer than 1000, 500, and 100 nt, respectively. For Velvet (k-mer size, coverage) we found that the coverage cutoff t = 5 maximizes the assembly quality. The effect of threading reads on assembly quality may be seen by comparing simBAC35 and simBAC100. The rows describing REPEAT-GRAPH(50) and OPTIMALASSEMBLY(50) are identical, since the reads cover the entire BAC in simBAC100 data set. In all tests there was a single misassembly (in simECOLI100 data set).











