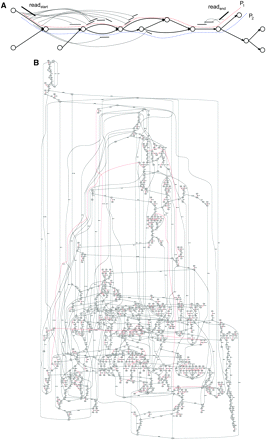
(A) A fragment of a made-up repeat graph formed by three divergent copies of a repeat. There are many possible paths from read start to read end . To transform the mate-pair, read star -GAP of length d-read end into a mate-read read start -SEQUENCE of length d-read end , we compute the support for every path between read start and read end and select a path with maximum support. In this example, the “red” path P 1 has greater support than the “blue” path P 2. (B) A fragment of the real repeat graph of E. coli (constructed from ECOLI data set) illustrating that transformation of mate-pairs into mate-reads may fail in some cases. Red edges represent unique (typically long) contigs, while black edges represent repeats.











