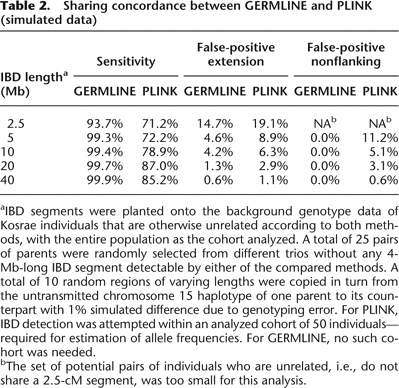
Sharing concordance between GERMLINE and PLINK (simulated data)
Click on table to view larger version.
-
aIBD segments were planted onto the background genotype data of Kosrae individuals that are otherwise unrelated according to both methods, with the entire population as the cohort analyzed. A total of 25 pairs of parents were randomly selected from different trios without any 4-Mb-long IBD segment detectable by either of the compared methods. A total of 10 random regions of varying lengths were copied in turn from the untransmitted chromosome 15 haplotype of one parent to its counterpart with 1% simulated difference due to genotyping error. For PLINK, IBD detection was attempted within an analyzed cohort of 50 individuals—required for estimation of allele frequencies. For GERMLINE, no such cohort was needed.
-
bThe set of potential pairs of individuals who are unrelated, i.e., do not share a 2.5-cM segment, was too small for this analysis.











