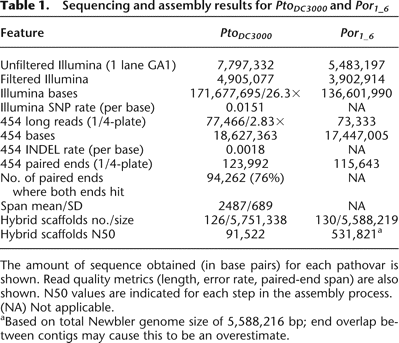
Table 1.
Sequencing and assembly results for PtoDC3000 and Por1_6
Click on table to view larger version.
-
The amount of sequence obtained (in base pairs) for each pathovar is shown. Read quality metrics (length, error rate, paired-end span) are also shown. N50 values are indicated for each step in the assembly process.
(NA) Not applicable.
-
aBased on total Newbler genome size of 5,588,216 bp; end overlap between contigs may cause this to be an overestimate.











