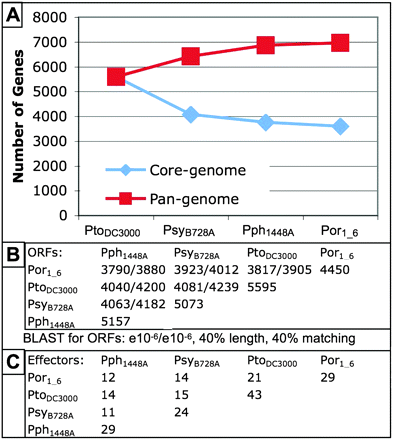
Por1_6 genome redefines the P. syringae pan-genome and type III effectorome. (A) The Por1_6 genome adds 97 unique genes to the pan-(distributed) genome of P. syringae (red boxes) and reduces the core (overlapping) genome by 161 genes (blue diamonds). (B) Por1_6 shares about 4000 of its 4450 predicted ORFs with each of the other three fully sequenced P. syringae strains. Shown are predictions based on a BLAST homology (E < 10−6). (C) Por1_6 contains a unique combination of type III effector proteins compared with the three previously sequenced strains. Type III effector proteins in Por1_6 were predicted using both homology with the known type III effectors in the other three fully sequenced P. syringae strains and gene prediction software (Methods). Note that the presence of a given type III effector gene, or homolog, does not necessarily imply that it is translocated (Chang et al. 2005; Schechter et al. 2006).











