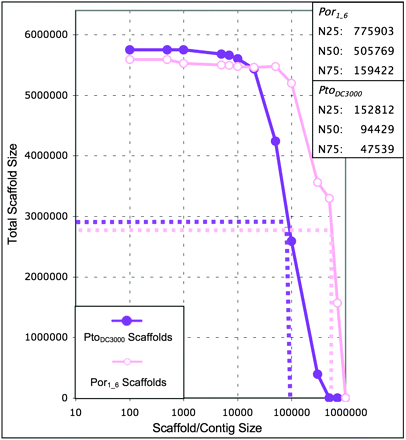
The Por1_6 genome assembled into contigs larger than 700,000 bp. Our hybrid assembly of Por1_6 is better than our hybrid assembly of PtoDC3000. The Por1_6 scaffolds (open pink circles) are significantly larger on average than the PtoDC3000 scaffolds (solid purple circles). The estimated N50 for Por1_6 is over 500,000 bp, with some scaffolds longer than 700,000 bp. N25, N50, and N75 values are estimated for Por1_6 based on total scaffold size (for Por1_6, 5,588,216 bp) rather than percentage of the genome covered as in Figure 3. This method is accurate: The PtoDC3000 N25, N50, and N75 values using this method are comparable to those derived from our syntenic approach (see Fig. 3).











