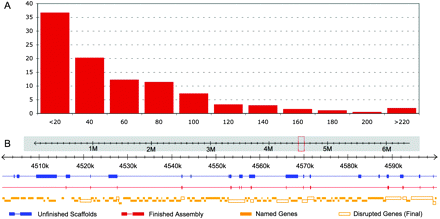
The final de novo PtoDC3000 hybrid assembly has few, mostly small gaps. (A) Histogram of gaps remaining in the PtoDC3000 hybrid assembly. Less than 1% of the reference genome (55,164 base pairs) remains unassembled into contigs or scaffolds. These bases are from a total of 1102 syntenic gaps with a median size of 55 bp. The largest gap is 987 bp (see also Supplemental data). (B) A typical 100-kbp region of the PtoDC3000 reference genome with ORFs shown (“Named Genes,” solid orange box) exhibits gaps remaining in the reassembled scaffold. “Unfinished Scaffolds” (blue) display the size and distribution of gaps before scaffold gaps are filled using unscaffolded contigs (Fig. 2D). Several large gaps are present. After finishing, most large gaps and many small gaps are eliminated (“Finished Scaffolds,” red). As a result, few (11/115) PtoDC3000 ORFs are disrupted (“Disrupted Genes,” open orange box) in this region.











