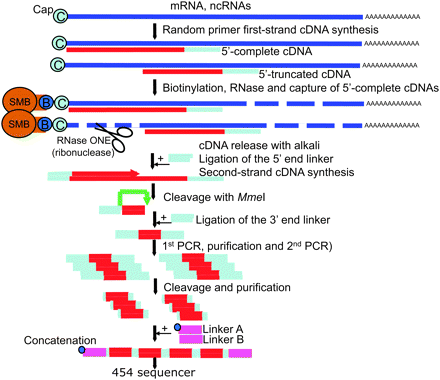
Preparation of DeepCAGE cDNA libraries. cDNA is produced with reverse transcriptase using random priming to maximize chances to reach the cap sites and to include non-polyadenylated RNAs. The cap site is biotinylated, followed by cleavage of single-strand RNA (incomplete cDNA). After recovery of the cDNA, a linker is ligated on the 5′-end, which carries the MmeI restriction site, which cleaves 20/21 nt inside the 5′-end of the cDNA. After a second linker ligation and PCR amplification, restricted digested tags are purified and finally concatenated together with the “454” linkers A and B. (SMB) Streptavidin-coated magnetic beads; (B) biotin.











