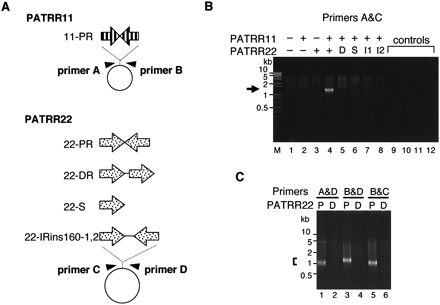
PCR detection of PATRR-mediated translocation-like rearrangements in HEK293 cells. (A) PATRR11 (11-PR, 445 bp) and PATRR22 (22-PR, 597 bp) were inserted into pUC19 and pBR322, respectively. Three modified versions of PATRR22—direct repeat (22-DR), single repeat unit (22-S), and inverted repeats with 156-bp spacer (22-IRins160-1 and 2)—were tested. Arrowheads indicate PCR primers for detection. (B) PCR specific for the translocation-like rearrangement. Plasmids (+) with or (−) without the PATRR were introduced simultaneously into HEK293 cells, and the joined molecules were detected by PCR. (Lanes 1–4, arrows) PCR products ∼1.2 kb in size originating from rearranged molecules were detected only when both of the PATRR-bearing plasmids were co-transfected. (Lanes 5–8) No such prominent PCR product was obtained from the modified versions of the PATRR22, (D) 22-DR, (S) 22-S, or (I1 and I2) 22-IRins160s. Smears and faint bands may be PCR artifacts possibly produced by nonspecifically degraded plasmids. DNA samples derived from (lane 9) cells without transfection, (lane 10) cells transfected with plasmids without addition of the transfection reagent, (lane 11) a simple mixture of two PATRR-bearing plasmids, and (lane 12) water without DNA were used as templates as negative controls. (Lane M) A 1-kb plus DNA ladder. (C) The rearrangement was also detected by PCR with three other combinations of primer pairs when the 11-PR plasmid was co-transfected with (P) 22-PR, but not with (D) 22-DR.











