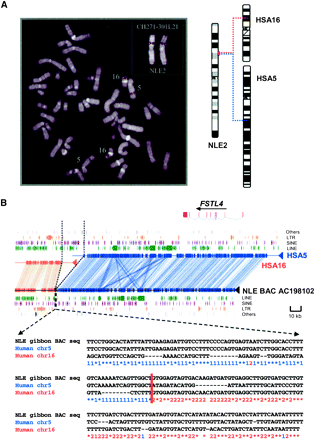
(A) Identification of gibbon BAC clones at the breakpoint of synteny. All BAC clones were experimentally validated by FISH as described previously (Roberto et al. 2007). In this example, a gibbon BAC clone spanning the breakpoint shows a single signal on gibbon chromosome 2 (NLE 2), but FISH mapped to human shows two signals on chromosomes 5 and 16, identifying an interchromosomal rearrangement (as represented by the chromosomal ideogram). (B) Sequence architecture at human–NLE gibbon synteny breaks. (Top panel) Miropeats analysis of the gibbon BAC, CH271-301L21 (AC198102), when compared with segments of human chromosome 5 (132461336–132644892, blue) and chromosome 16 (73369800–73421145, orange). Representative repeat elements, LINE, SINEs, LTRs, segmental duplications, and genes mapping to the synteny blocks with arrows denoting transcriptional orientation are also shown based on human genome annotation. (Bottom panel) Three-way ClustalW alignment between human and NLE gibbon sequences at the breakpoint with 1 (blue) denoting a sequence identity with the human chromosome 5 segment and 2 (orange) indicating sequence identity with the human chromosome 16 segment. The figure shows a class I breakpoint where the human breakpoints abut precisely at the point of fusion on the gibbon chromosome.











