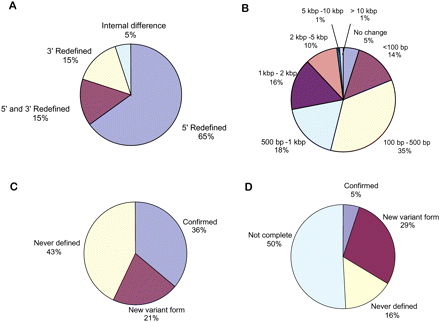
Generated transcript and ORF models compared with the corresponding WormBase annotation. (A) The percentage of new ORF models that differ from the corresponding WormBase models. If an ORF model had internal changes coexisting with changes in boundary (redefined 5′-, 3′-, or both ends), the ORF model was counted as having a change in boundary. (B) Chromosomal span change of novel ORF models in kilobases (kb). A change can be either an increase or a decrease in span. (C,D) Changes in annotated UTRs. UTRs in the RD-transcript models are divided into several categories: confirmed (WormBase), new variant (different from WormBase), and never previously defined (no UTR indicated in WormBase). Many transcript models in WormBase do not have UTRs defined, and a model can be annotated as having multiple UTRs. “Not complete” UTRs are those 3′ UTRs that could not be defined with certainty.











